Key Engineering Contributions
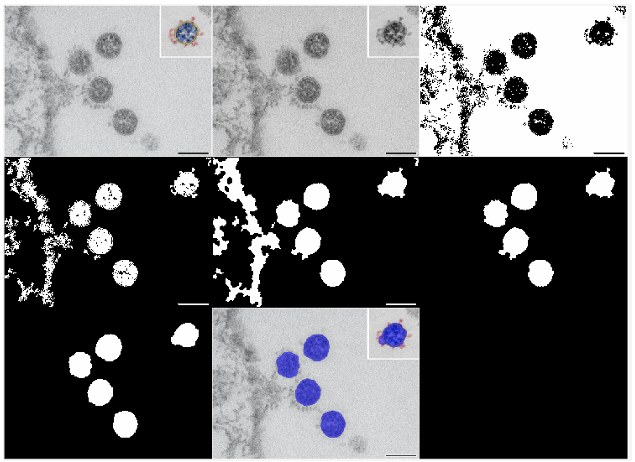
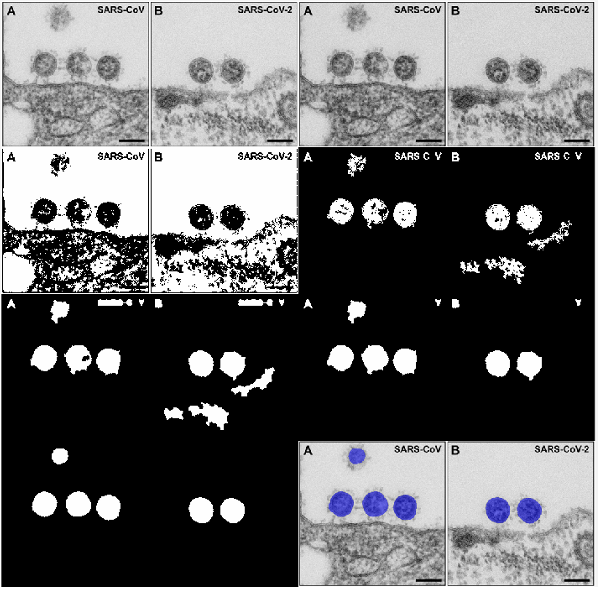
- 01Image Segmentation: Implemented Otsu's method to binarize grayscale TEM images, successfully isolating the viral particles from the background cellular environment.
- 02Morphological Filtering: Applied custom morphological closing operations and strict geometric thresholds (area boundaries and eccentricity ≤ 0.75) to automatically filter out pixel noise, artifacts, and incomplete cell boundaries.
- 03Quantitative Data Analysis: Calculated and converted the pixel-based area of the identified cells into physical dimensions, successfully extracting the centroid coordinates and determining a cumulative viral area of 41,151 nm2 across the analyzed sample.
Visual Documentation